The Pillai lab identified that NUDT12 is a decapping enzyme of NAD-capped RNAs in the cytoplasm and forms a complex with BLMH, which is responsible for the localization of the complex. Their findings were published in Cell Reports.
Highlights
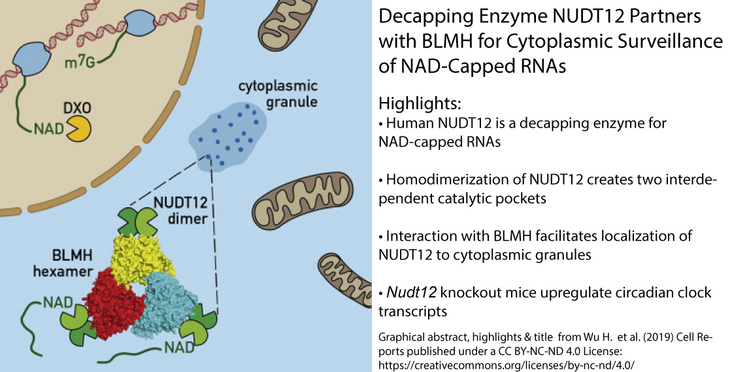
- Human NUDT12 is a decapping enzyme for NAD-capped RNAs
- Homodimerization of NUDT12 creates two interdependent catalytic pockets
- Interaction with BLMH facilitates localization of NUDT12 to cytoplasmic granules
- Nudt12 knockout mice upregulate circadian clock transcripts
Summary
RNA polymerase II transcripts receive a protective 5',5'-triphosphate-linked 7-methylguanosine (m7G) cap, and its removal by decapping enzymes like DCP2 is critical for initiation of RNA decay. Alternative RNA caps can be acquired when transcription initiation uses metabolites like nicotinamide adenine dinucleotide (NAD), generating NAD-RNAs. Here, we identify human NUDT12 as a cytosolic NAD-RNA decapping enzyme. NUDT12 is active only as homodimers, with each monomer contributing to creation of the two functional catalytic pockets. We identify an ~600-kDa dodecamer complex between bleomycin hydrolase (BLMH) and NUDT12, with BLMH being required for localization of NUDT12 to a few discrete cytoplasmic granules that are distinct from P-bodies. Both proteins downregulate gene expression when artificially tethered to a reporter RNA in vivo. Furthermore, loss of Nudt12 results in a significant upregulation of circadian clock transcripts in mouse liver. Overall, our study points to a physiological role for NUDT12 in the cytosolic surveillance of NAD-RNAs.
Read the Publication in Cell Reports (Open Access)
Graphical abstract, highlights, summary & title from Wu H. et al. (2019) Cell Reports published under the CC BY-NC-ND 4.0 License.