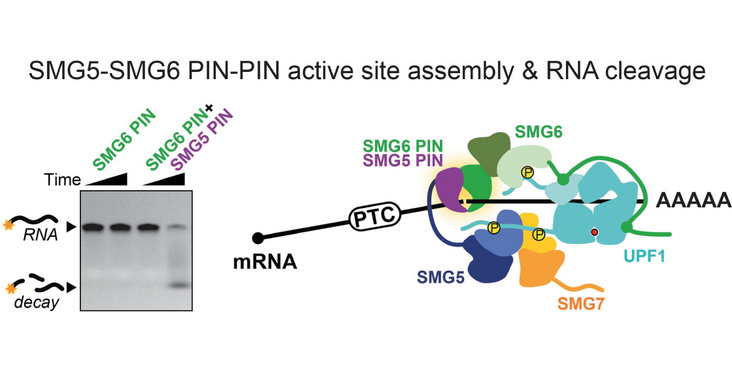
The Gatfield (University of Lausanne) and Jonas (ETH Zürich) labs discovered that both PIN nuclease domains of SMG5 and SMG6 are required for efficient mRNA degradation by Nonsense-mediated decay. The PIN domain of SMG5 was considered to not be involved in mRNA degradation. Now, the researchers could show that the PIN domain of SMG5 contributes an aspartate residue to the acidic triad of the SMG6 PIN domain thereby creating the canonical acidic tetrad of PIN domains. Their findings have been published in the article “Active Site Assembly by SMG5 as a Mechanism for SMG6 Endonuclease Licencing in Nonsense-mediated mRNA Decay” in the Journal of Molecular Biology.
Highlights:
- SMG5-SMG6 PIN domains form a conserved composite nuclease centre.
- SMG5 provides catalytic residues that mechanistically explain SMG6 licencing.
- SMG5 and SMG6 extensions stabilise a weak, SMG5-SMG6 PIN-PIN interface.
- Covalent tethering restores activity by enforcing SMG5-SMG6 proximity.
- Real?time fluorescence assay quantifies SMG5?dependent SMG6 activation.
Abstract:
Nonsense-mediated mRNA decay (NMD) is a conserved eukaryotic surveillance pathway that eliminates transcripts containing premature termination codons (PTCs). Substantial progress has been made in defining the transcript features that mark aberrant translation termination for NMD activation, yet key mechanistic steps remain incompletely understood – including how recruitment of the central NMD factor UPF1 is coupled to the downstream effector phase in which targeted mRNAs are nucleolytically degraded. In metazoans, NMD employs an endonucleolytic route mediated by SMG6, a PIN-domain nuclease, alongside SMG5 and SMG7, which act downstream of PTC recognition. SMG5 has recently been proposed to licence SMG6 activity, yet the molecular basis of this licencing has remained elusive. Here, we combine AlphaFold structural predictions with biochemical assays to investigate interactions among human SMG5, SMG6, and SMG7. Structural models predict a high-confidence interface between SMG5 and SMG6 PIN domains that forms a composite active site: a conserved SMG5 aspartate (D893) complements the SMG6 acidic triad to reinstate the canonical tetrad required for PIN-domain catalysis. In vitro, SMG6 alone exhibits weak endonucleolytic activity, which is enhanced ~10-fold by the SMG5 PIN domain. Mutational analyses confirm that conserved residues from both proteins are essential for this composite configuration. Our findings reveal that the SMG5 PIN domain, previously considered catalytically inert, plays a critical role in activating SMG6 by completing its active site. This work provides mechanistic insight into the SMG5-dependent licencing step and uncovers a composite PIN nuclease architecture at the heart of the metazoan NMD effector phase.
Read the Publication in the Journal of Molecular Biology (Open Access)
Abstract, figure and highlights from Arpa, Taschner, et al. (2026) J Mol Biol published under a CC BY 4.0 license.