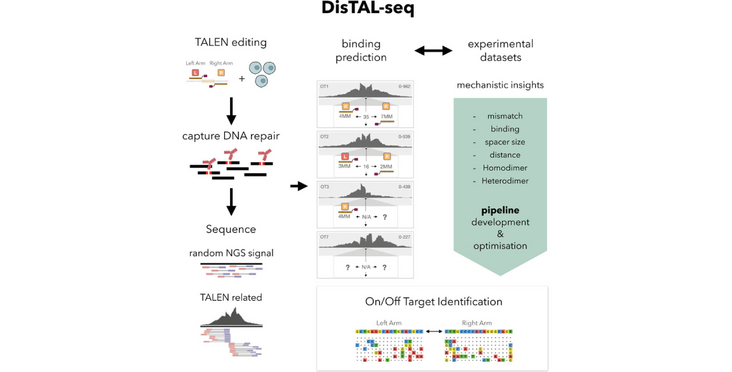
Despite the widespread use of CRISPR-based genome editing nucleases in basic research and the clinic, Transcription Activator-Like Effector Nucleases (TALENs) are being applied in the clinic, and relatively little is known about their target recognition and specificity. The Corn lab and collaborators now adapted their DISCOVER-Seq pipeline to detect CRISPR off-targets to TALENs. The adapted pipeline called DisTal-Seq was applied to different TALENs and T cell donors and gave insights into the specificity of the respective TALEN. The pipeline should help the development of safer and more effective TALENs and was described in the article “DisTAL-Seq: A TALEN-specific adaptation of DISCOVER-Seq for off-target profiling” in Molecular Therapy Nucleic Acids.
Abstract
Programmable guided nucleases have revolutionized genome editing and biomedical research, with transformative potential for gene and cell therapy. Although the widespread adoption of the CRISPR-Cas system has provided deep insights into target recognition and specificity, the behavior of clinically relevant tools like transcription activator-like effector nucleases (TALENs) remains poorly characterized in human cells. To address this gap, we implemented DisTAL-Seq, a TALEN-specific adaptation of the DISCOVER-Seq pipeline, which detects MRE11 recruitment to double-strand breaks (DSBs). Based on the DISCOVER-Seq principle, DisTAL-Seq incorporates alignment logic tailored to TALEN-binding properties, including variable RVD specificity, cleavage offset, and dimerization behavior. Using DisTAL-Seq, we identified and validated on- and off-target sites across diverse TALENs and T cell donors. This unbiased approach revealed key features of TALEN activity in human cells, including number of tolerated mismatches to a target site and relative location of the induced DSB. DisTAL-Seq thus extends DISCOVER-Seq to the TALEN family and provides a robust platform for assessing modifications in enzyme architecture and application contexts on a genome-wide scale, supporting the development of safer and more effective genome editing tools.
Read the Publication in Molecular Therapy Nucleic Acids (Open Access)
Abstract, figure and title from Kobel et al (2026) Mol Ther Nucleic Acids published under a CC BY 4.0 license.