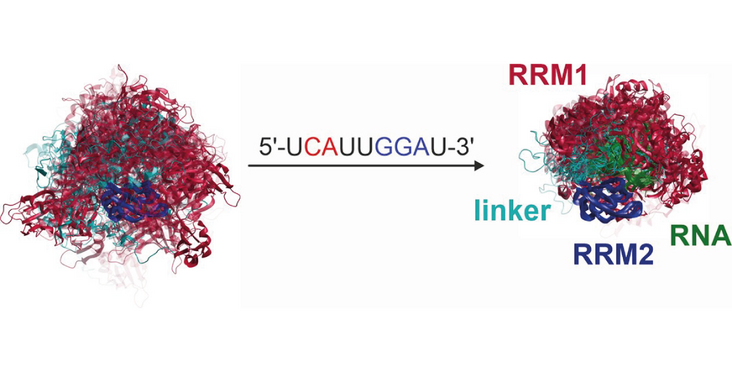
Biomolecular binding approaches can be much more dynamic as some structural snapshots, especially of components, suggest. The Allain and Jeschke labs using an integrative approach could characterize the dynamic binding conformations of the tandem RRMs of SRSF1, which are separated by a flexible linker, to RNAs, which result in a different orientation of the RRMs of the RRM. They used NMR paramagnetic relaxation enhancement (PRE) and electron paramagnetic resonance (EPR) distribution restraints. The conform ensembles showed clear difference between the free and the bound state, as well as depending on the binding orientation of the RRMs. AlphaFold3 based approaches proved to be less successful in generating ensembles, but the authors could derive a less demanding approach which only requires PRE data and the use of a different computational application. Their findings have been published in the article “Characterization of flexible RNA binding by tandem RNA recognition motifs through integrative ensemble modelling ” in the journal Nucleic Acids Research.
Abstract
Multi-functional, promiscuous RNA-binding proteins may exhibit substantial disorder in both their free and bound states. The tandem RNA recognition motifs (RRMs) of SRSF1 can serve as a model for such behaviour, as they bind RNA with RRM1 either upstream or downstream from RRM2. Here, we study binding of two short single-stranded RNAs with different directionality to a construct featuring the tandem RRMs and their 30-residue flexible linker. We integrate NMR paramagnetic relaxation enhancement (PRE) and electron paramagnetic resonance (EPR) distance distribution restraints for ensemble modelling. Binding only moderately reduces the occupied conformation space. Ensembles with RRM2 bound downstream or upstream from RRM1 are clearly distinguishable. The system can serve as a testbed for integrative ensemble modelling approaches. Initial conformer pools generated by AlphaFold3 are more compact than those generated by other approaches. AlphaFold3 fails to predict RNA binding of RRM2. Ensembles generated with MMMx using rigid body search to fit distance distributions, followed by selection of a population-weighed subset of conformers consistent with both EPR data and PREs, gave the best fit, and were robust to elimination of individual distance distributions. We also present a less demanding PRE-only approach using multi-state structure calculations with CYANA.
Read the Article in Nucleic Acids Research (Open Access)
Abstract, figure and title from Nguyen et al (2026) Nucleic Acids Res published under a CC BY 4.0 license.