Ribosomal ubiquitin-like fusion protein: Same same but different?
In most eukaryotes, one ribosomal protein of each subunit is initially translated as an in cis linear fusion with a ubiquitin moiety. These N-terminal ubiquitin moieties are cleaved off during ribosome biogenesis. The cleaved-off ubiquitin is a major source of cellular ubiquitin, which can be conjugated to other proteins. It can only be speculated about why such a link between ribosome biogenesis and the ubiquitin-based protein degradation system exists. It could serve to coordinate the cellular protein synthesis rate with degradation.
Another type of ribosomal fusion protein, FUBI-eS30, exists in humans and other holozoan organisms. The N-terminal half of this fusion protein consists of the ubiquitin-like protein FUBI, while the C-terminal eS30 is a ribosomal protein being part of the mature 40S subunit. The Kutay lab, which has a long-standing interest in ribosome biogenesis, set out to investigate FUBI-eS30.
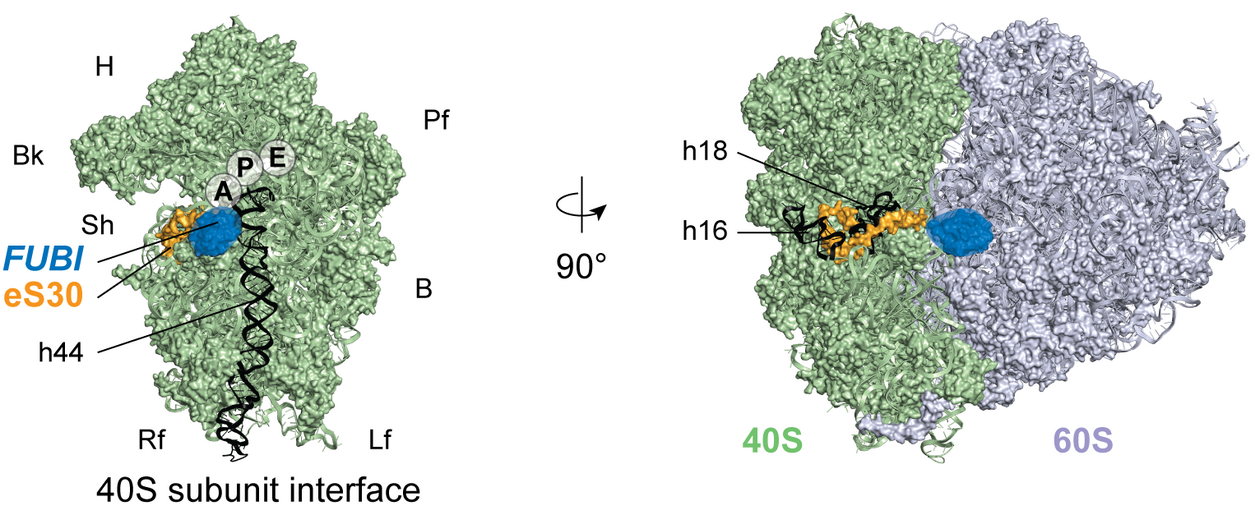
For the classical ribosomal ubiquitin fusion proteins, it is known that only the ribosomal protein parts of the fusion proteins, but not ubiquitin, are part of mature ribosomes. This becomes already apparent when looking at ribosome structures because uncleaved ubiquitin moieties would sterically impair the ribosome function. The same holds true for FUBI. “When working on ribosome biogenesis, I enjoyed the availability of detailed structural data. It gives you a visual idea of what you are working with and helps to view your hypotheses from different angles,” says Jasmin van den Heuvel, first author of the study published in eLife and former PhD student in the Kutay lab.
Like ubiquitin, FUBI contains at its C-terminus a double glycine motif. Mutation of this motif hinders cleavage of the ubiquitin moiety from its in cis fusion protein. The researchers created two FUBI-eS30 mutants based on previous literature in which the diglycine motif was mutated. These constructs also contained a tandem StHA tag at the C-terminus of eS30, and as a control, a wild-type construct with the same C-terminal tag was created. Compared to the wild-type construct, the diglycine mutant proteins were not cleaved in cells. Also, their expression led to decreased 40S and polysome levels. At the same time, increased 60S levels were observed, which was previously reported when 40S production was hindered. Through further experiments, they could dissect that the non-cleavable mutants led to a defect in the late 40S maturation in the cytoplasm.
The researchers performed mass spectrometry with the previously mentioned constructs to identify the protease responsible for cleaving FUBI from eS30. Two known deubiquitinases (DUBs) were enriched in the pull-downs of the uncleavable constructs compared to the wild-type construct. Depletion experiments with siRNAs showed that only one of them, USP36, was responsible for cellular FUBI cleavage. Using CRISPRi for the depletion of endogenous USP36, only a wild type USP36 containing construct, but not one with a catalytically inactive mutant, could revert the accumulation of uncleaved FUBI-eS30 in cells. Also, they showed that USP36 cleaves FUBI-eS30 in vitro. Based on these experiments, USP36 is a bispecific enzyme that can cleave ubiquitin and FUBI off its fusion protein.
Like for ribosomal ubiquitin fusion proteins, the FUBI moiety of the fusion protein with the ribosomal protein eS30 needs to be cleaved by a DUB to create functional ribosomes. An open question is if FUBI, like ubiquitin and some other ubiquitin-like proteins, is used to modify other proteins. Although it cannot be ruled out that other enzymes are involved in the cleavage of FUBI-eS30, USP36 seems to be the key player in this process. Interestingly, the oncogenic transcription factor Myc is one of the known proteins that USP36 deubiquitinates and thereby stabilizes. Thus, USP36 could be an interesting target for cancer drug development. “I was initially skeptical whether anti-cancer drugs can be used to target fundamental processes such as ribosome biogenesis or nuclear transport without killing too many healthy cells in an organism. But the first drugs targeting these processes seem to show that at low doses, cancer cells are susceptible to these drugs while normal cells can tolerate such doses,” says Ulrike Kutay, last author of the study and Full Professor at the Institute of Biochemistry at ETH Zurich.
Publication: van den Heuvel et al. (2021) eLife, 10, e70560 (Open Access)
Figure from van den Heuvel et al. (2021) eLife published under a CC BY 4.0 license.
Text: Dominik Theler