SARS-CoV-2 depends on the ribosome missing a step
Many viruses depend on ribosomal frameshifting for the expression of their proteome and replication. Such viruses include HIV, Influenza A as well as all known Coronaviruses. Coronaviruses require a -1 frameshift so that its open reading frame (ORF) 1b in addition to the ORF1a can be translated. ORF1b encodes for the RNA-dependent RNA polymerase and additional proteins crucial for viral replication. Although frameshifting is critically important for the replication of medically relevant viruses, so far, no detailed molecular image of the process could be obtained.
To address this question, the Atkins (University of Cork, Ireland) and the Ban labs (ETH Zurich) teamed up and were later joined by the Bode (ETH Zurich), Gatfield (University of Lausanne) and Thiel (University of Bern) groups. Initially, the researchers established the biochemistry to trap the ribosome in order to investigate a cellular frameshifting event. “When the Covid-19 pandemic broke out, this previous research allowed us to rapidly switch to study the frameshifting site of the SARS-CoV-2 virus.” says Alain Scaiola, co-first author of the study published in Science and PhD student in the Ban lab.
By Cryo-EM, they solved the structure of the 80S ribosome paused at the SARS-CoV-2 frameshifting site. For this complex, the core of the ribosome was resolved at 2.2.-Å resolution, which is the highest resolved structure of a mammalian ribosome to date. Consequently, as an extra bonus of the study, the high-resolution information allowed for the visualization of many protein and rRNA modifications never directly seen before but known from other methods. “We were stunned by the resolution obtained since we did not push for this methodologically. The resolution results from the ribosome adopting a strained conformation when trapped at the frameshifting site.” explains Scaiola.
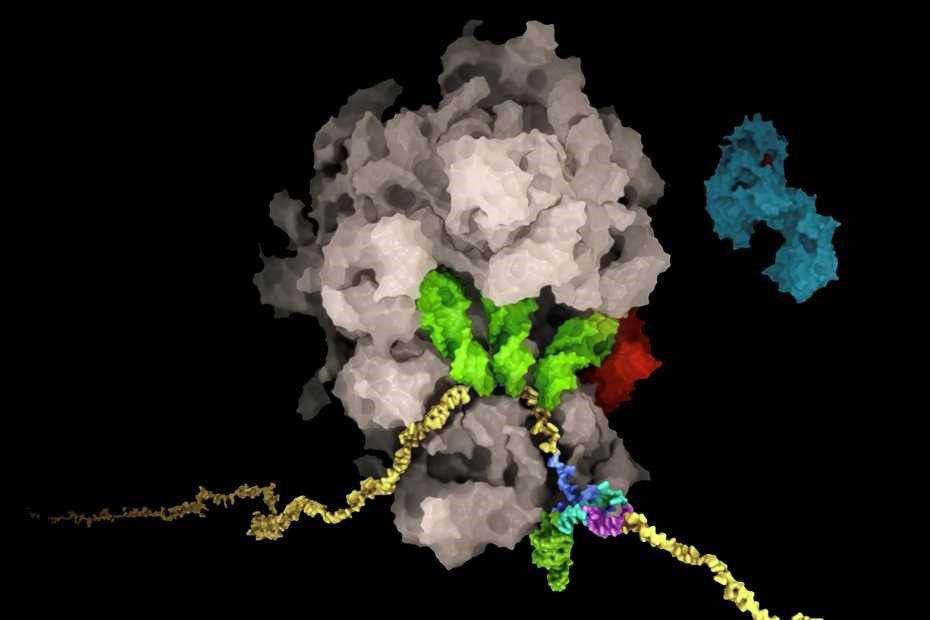
From previous research, it was known that the frameshifting occurs only when the ribosome interacts with the slippery frameshifting site and as well with a pseudoknot structure downstream. These interactions could be visualized in the structure. To the big surprise of the researchers, specific interactions also take place between the ribosomal exit tunnel and the nascent chain being translated when the ribosome hits the frameshifting site. Two of the amino acids of the nascent chain that interact with the ribosome show a high degree of conservation among Coronaviruses, despite their location in an unstructured region of the viral protein.
Based on their structure, the researchers performed follow-up biochemical and cellular experiments. With the combination of the applied experimental approaches, the researchers obtained a detailed molecular and mechanistic understanding of this intricate process essential for Coronavirus replication. So far, similar ribosomal frameshifting has not been reported for native cellular transcripts, so that its inhibition by drugs could be a way to treat all types of Coronavirus infections. “Based on how conserved this process is, as well as the interactions between the ribosome, the viral RNA and the nascent chain, it would be difficult for the viruses to evolve mutations, making them resistant to drugs targeting the frameshifting event.” says Scaiola.
In the literature, two compounds targeting the pseudoknot were proposed to inhibit the frameshifting, and only one of them was tested in infected cells. When the researchers treated SARS-CoV-2 infected African green monkey cells with the two compounds, viral replication was decreased up to four orders of magnitude without apparent cellular toxicity caused by the compounds. Interestingly, contrary to previous reports, only one of the two had a dose-dependent effect on frameshifting, while the other seems to reduce viral replication through a different mechanism. Although these compounds are not potent enough for being used as drugs, the results provide a starting point for drug development.
Bhatt, Scaiola et al. (2021) Science 372(6548), 1306-1313 (Open Access)
Text: Dominik Theler