It matters how we (or RNA) are born
561-569 (2017).
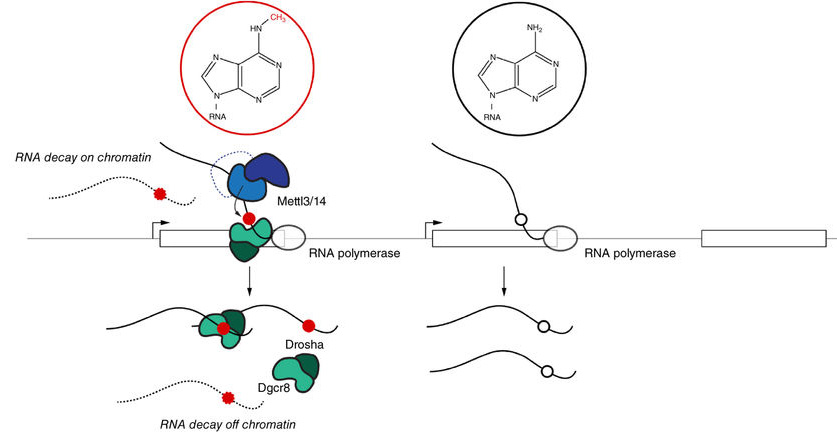
Eukaryotic gene expression is heavily regulated at the transcriptional and posttranscriptional levels. An additional layer of regulation occurs co-transcriptionally through processing and decay of nascent transcripts physically associated with chromatin. This process involves components of the RNA interference (RNAi) machinery and is well documented in yeast, but little is known about its conservation in mammals. Yet it is clear that control of RNA lifespan is vital for the proper functioning of our cells. Marc Bühler's group at the Friedrich Miescher Institute for Biomedical Research (FMI) in Basel has discovered a novel mechanism determining the fate of RNA in mammalian cells: two proteins of the RNAi machinery- Dgcr8 and Drosha - together with a methyltransferase, Mettl3, mark nascent RNAs for degradation right in the moment they are transcribed.
The formation, processing and degradation of RNA are all tightly regulated. This stringent control of RNA metabolism ensures that genes become active at the right time and place, making sure cell functions are always well orchestrated. In this context, control mechanisms collectively referred to as RNA interference (RNAi) has attracted a lot of attention. Commonly, RNAi leads to the fragmentation and inactivation of RNAs in the cytoplasm. Interestingly, in yeast, an active RNAi machinery can also be found in the nucleus: during RNA synthesis, while the RNA molecules still associate with the DNA, it triggers the prompt degradation of nascent RNAs.
Marc Bühler and his group at the FMI used mouse embryonic stem cells to find out whether the RNAi machinery might play a similar role in mammalian cells. Bühler comments: "This is a good example of how knowledge gained in a model organism - here in fission yeast - guides our hypotheses and informs our experiments in higher organisms." His group's previous work in S. pombe revealed that RNAi as well as other nuclear RNA degradation pathways mediate the degradation of RNA in association with chromatin. But is such regulation evolutionarily conserved? Yes, they were indeed able to find a similar mechanism in mammals, albeit with major differences to the yeast system. For example, Dicer does not physically associate with chromatin. Instead, their results suggest that the Microprocessor (con-sisting of Dgcr8 and Drosha), which does not exist in yeast, takes over this function in multicellular organisms. The FMI scientists also showed that an enzyme known as Mettl3 is involved in the degradation of nascent RNAs. Mettl3 transfers methyl groups to adenosine residues in RNA, a mark that also influences RNA stability.
The Microprocessor/Mettl3 system allows the cell to react rapidly to changing growth conditions. Interestingly, Bühler's group has found that the novel mechanism helps cells to cope with heat stress. There, genetic ablation of Dgcr8 or Mettl3 leads to the accumulation of Hsp70 mRNA, elongation of its half-life, and an increase in protein levels - so they propose that acute heat stress co-transcriptionally marks Hsp70 mRNAs for subsequent RNA degradation to control the timing and magnitude of the heat-shock response. According to Bühler, both the fast stress response and the rapid clearance of heat-shock transcripts and proteins are important for cells: "The accumulation of stress response proteins is detrimental to the cell and is often observed in cancer." The new findings shed light on the important role of co-transcriptional regulation of genes, especially in such full-speed/full-stop cases. In fact, Bühler would not be surprised if many more of such co-transcriptional markings would be found - if the RNA remembered a lot more details from the moment of its birth than is actually known.
Knuckles P. et al. (2017) Nature Structural & Molecular Biology 24, 561-569
By Roland Fischer