miRNAs caught in the act
This must be a bit like colonizing a new land, for the molecular biologist. But pioneers need maps. They need to know how things are connected. They need to know which paths lead where.
A multidisciplinary initiative from the ETH Zürich and the Biozentrum at the University of Basel, amongst others, may help finding the way. Most biological processes in mammals are regulated by miRNAs, and identification of miRNA targets therefore is one of the big challenges of molecular biology. Complex target site prediction algorithms have been very successful in matching the 6- to 8-nucleotide “seed” region of a miRNA to potential targets, but they do not provide enough information to reliably predict unorthodox miRNA-RNA interactions. Now Imig and Brunschweiger et al. figured out a way to capture miRNA targets in cells biochemically. Their approach, termed miR- NA crosslinking and immuno- precipitation (miR-CLIP), uses pre-miRNAs doubly modified with photoreactive psoralen and biotin groups.
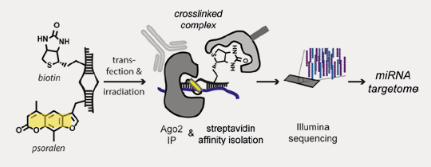
MiR-CLIP isolates miRNA targets using a multistep process. After delivery of the synthetic miRNA, RNA-protein crosslinks are induced within the silencing complex, and then the psoralen is activated, forming crosslinks between the miRNA and target RNA. The crosslinked complex, containing synthetic miRNA, target RNA and protein components of the silencing machinery, is then isolated in a two-step process by immunoprecipitation against Ago2, a silencing complex component and then streptavidin affinity purification.
MiR-CLIP was established in HeLa cells, seeking targets of miR-106a. Many of the iden- tified targets corresponded to canonical targets predicted by predictive tools. Furthermore miR- CLIP identified numerous new targets, though the validity of these putative targets is not yet clear. “miRNAs will usually interact with hun- dreds or even thousands of sites in the cell”, the group leader Jonathan Hall from the ETH Zürich explains. “To be able to take a snapshot and to capture these interactions may very well open up new avenues of research.”
Indeed, one unexpected and interesting find was that H19, a large intergenic noncoding RNA (lincRNA), was identified as a target of miR-106a. LincRNAs are a diverse and mysterious class of long noncoding transcripts important for gene regulation.
Together, these studies suggest that there are yet-undiscovered mechanisms of post-transcriptional regulation through miRNAs. And, as the authors conclude: “They also highlight a need for new experimental methods to complement and expand existing computational target prediction methods.”
By Roland Fischer