Clogging the Ribosome to go Viral
While the entire world is on alert due to the current COVID-19 pandemic, scientists are seeking to understand the molecular mechanisms underlying the virulence of SARS-CoV-2 in order to rapidly develop drugs and vaccines against COVID-19. A collaboration among three labs from the NCCR RNA & Disease network (Ban laboratory at ETH Zurich and Mühlemann and Thiel laboratories at University of Bern) has now resulted in a publication in Nature Structural & Molecular Biology, in which they describe how the SARS-CoV-2 virus hijacks the host cell to promote its own replication (Schubert, Karousis et al., 2020).
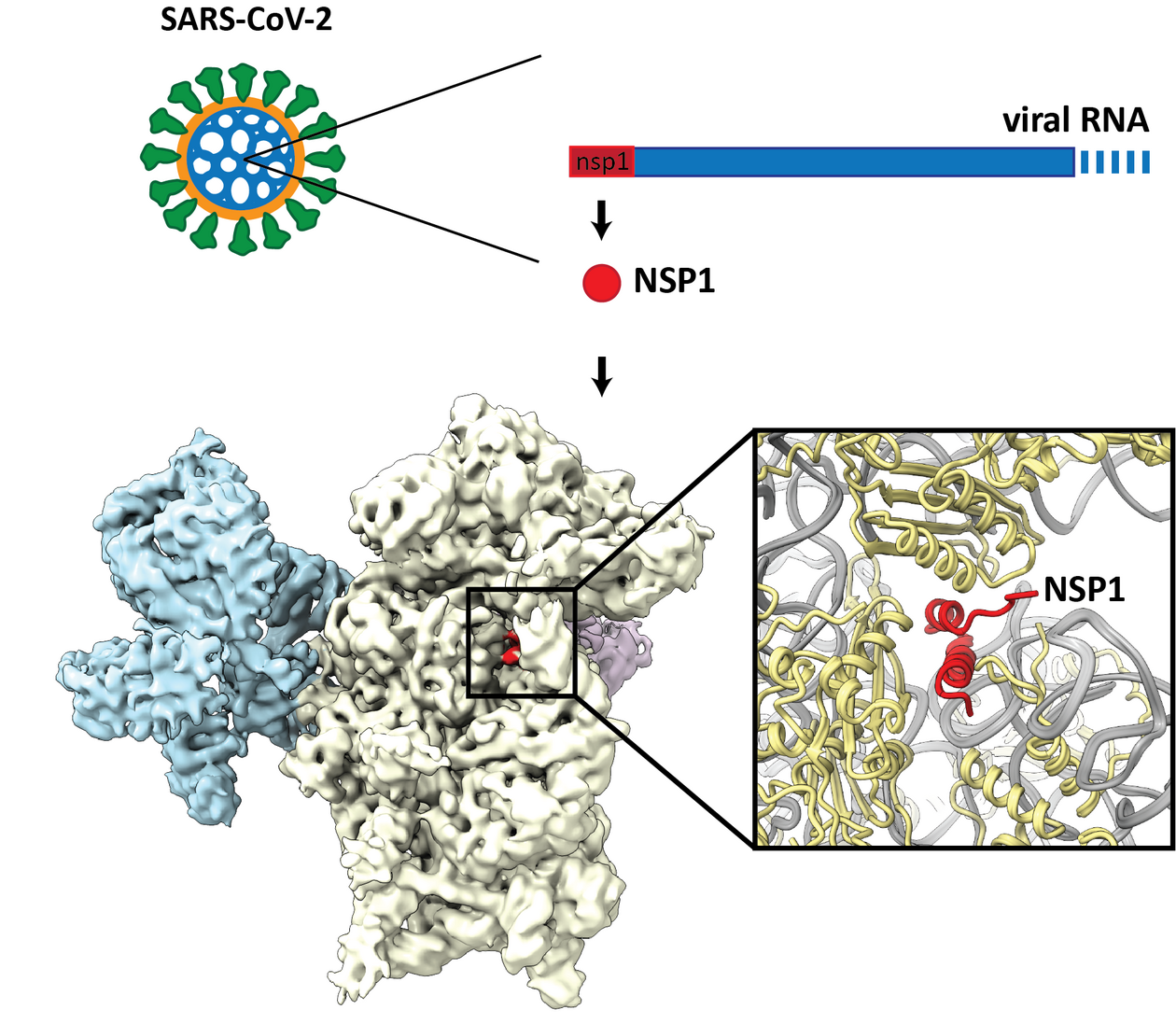
Upon host cell invasion, one of the first viral proteins translated is the “non-structural protein 1” (Nsp1). Despite its unspectacular name, Nsp1 is a major virulence factor of certain coronaviruses including SARS-CoV-1 and -2 and has been proposed to downregulate the host innate immune response to generate a virus-friendly cellular environment for viral replication. On the molecular level, Nsp1 associates with the ribosome and represses host gene expression, but the underlying molecular mechanism remained enigmatic.
To explore the mechanism of how Nsp1 alters host translation, Schubert, Karousis and colleagues set out to solve the structure of Nsp1 interacting with the ribosomal complexes using cryo-EM. The structure already hinted towards a potential mechanism of how Nsp1 inhibits translation: Part of the Nsp1 protein binds to the site of the ribosome where mRNAs normally enter during translation initiation. Together with the fact that RNA substrates were not observed in the Nsp1-ribosome structures, this suggests that Nsp1 inhibits translation by acting as a steric roadblock for mRNAs, preventing its association with the translation machinery.
The functional consequence of the Nsp1-ribosome interaction was further examined in an in vitro translation assay, which was recently developed in the Mühlemann laboratory and has recently been published in Nature Communications (Karousis et al., 2020). This in vitro translation system allows to monitor the ribosome occupancy on reporter mRNAs in mammalian cell lysates. With this tool in hands, the authors could validate their hypothesis that Nsp1 can effectively inhibit translation of a reporter mRNA. The detailed structure of Nsp1 allowed the design of Nsp1 mutants that are not able to bind to the 40S ribosomal subunit in vitro. These Nsp1 mutants could no longer inhibit translation both in vitro as shown in the translation assay and in vivo as shown in a puromycin incorporation experiment in HeLa cells.
If the virus inhibits the ribosome, how can the virus itself evade Nsp1-mediated translation inhibition to ensure the translation of its own messages? To solve this conundrum, the authors modified the reporter mRNA by replacing the 5'UTR of the reporter mRNA with the 5'UTR of the SARS-CoV-2 genomic RNA. They compared the translation efficiency of these transcripts using the in vitro translation assay and observed that the reporter mRNA harbouring the viral sequence is more efficiently translated than the native reporter transcript. When Nsp1 is added to the reaction, both reporter mRNAs are affected by the translation inhibition to the same extent. Oliver Mühlemann, Professor of Biochemistry at the University of Bern and one of the co-corresponding authors of the study, explains a potential mechanism of how viral mRNAs could be translated efficiently despite the global shut-down of ribosomes: “When Nsp1 blocks the ribosomes, functional ribosomes become scarce and at the same time viral RNA can make up almost half of the total RNA in the cell. Under these conditions, the viral RNA is preferentially read from the still functional ribosomes compared to the cell's own messenger RNA.” The authors further speculate that this mechanism also suppresses host mRNAs required for innate immune responses. The combination of these effects could provide an ideal cellular environment for viral replication and contribute to the successful replication and dissemination of SARS-CoV-2.
Due to its essential role in promoting virulence to SARS-CoV-2, Nsp1 represents an attractive target for future drug and attenuated vaccine development. Nenad Ban, Professor for Molecular Biology at ETH Zurich and co-corresponding author of the study, explains the importance of the published results: “The high-resolution structure of Nsp1 interacting with the ribosome provides important information for the design of a drug that could prevent Nsp1 from binding to the ribosome without interfering with the ribosomal function. If Nsp1 can no longer inhibit translation, we expect that the cellular defence system can be active and stop the viral replication.”
Schubert & Karousis et al. (2020) Nat Struct Mol Biol, 27(10), 959-966
Karousis et al. (2020) Nat Commun, 11(1), 4134
Text: Veronika Herzog