Ribosomal proteins taking over
In a recently published Science paper, David J. F. Ramrath and Moritz Niemann from the Ban and Schneider groups shed light on the evolutionary shift toward protein-based architecture in trypanosomal mitochondrial ribosomes.
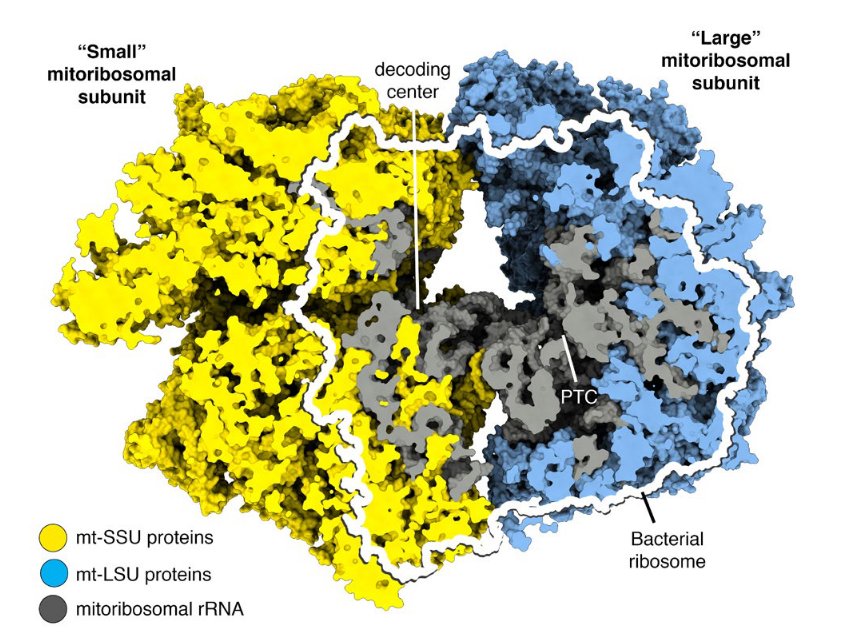
Mitochondrial ribosomes (mitoribosomes) are more closely related to bacterial ribosomes than to eukaryotic cytosolic ribosomes. However, they have undergone extensive structural and compositional change throughout evolutionary time. Most notably, mitoribosomes have acquired a large number of mi- tochondrialspecific ribosomal proteins, and the mitoribosomal RNA has been shortened in many organisms, including mammals, but most extensively in Trypanosoma brucei, the parasite that causes sleeping sickness.
Because mitoribosomes are conserved to a high degree, the observed variability is of particular interest, especially in extreme cases as T. brucei. In these mitoribosomes featuring the smallest known rRNAs, the severe rRNA reduction is accompanied by the recruitment of many additional proteins. Trypanosomal mitoribosomes therefore represent an excellent system to reveal the minimal set of rRNA and protein elements essential for ribosomal function and to investigate how ribosomal proteins compensated for the missing rRNA.
To address these questions, the Ban and Schneider groups determined the atomic structure of the mitoribosome from T. brucei using cryoelectron microscopy. The structure shows how the proteins have taken over the role of architectural scaffold from the rRNA: They form anautonomous outer shell that surrounds the entire particle and stabilizes the functionally important regions of the rRNA. The paper reveals the “minimal” set of conserved rRNA and protein components shared by all ribosomes, which will help define the most essential functional elements.
The trypanosomal translational machinery adopted unusual solutions to accomplish some basic protein synthesis mechanisms. Notably, the structure unveils two intriguing functional details: Their nascent polypeptide exit tunnel branches into two exits – it is conceivable that nascent proteins with different characteristics take different paths. Furthermore, in a subpopulation of isolated small-subunit particles, mitochondrial initiation factor 3 was observed interacting with the decoding center via its unique C-terminal extension. This might compensate for the essential function of initiation factor 1 that is absent in all mitochondria.
Ramrath and Niemann et.al., (2018) Science 362, eaau7735
By Roland Fischer