Through the use of a conditional Smg 6 mutant, which is a key protein for the Non_sense Mediated Decay (NMD) system, the Gatfield lab and collaborators could show that NMD is involved in the regulation of the circadian clock. Thus, showing that NMD is involved in regulating vivo in a physilogical not coupled to the degradation of erroreneous transcripts. Their findings have been published in the article "A conditional Smg6 mutant mouse model reveals circadian clock regulation through the nonsense-mediated mRNA decay pathway" in Science Advances.
Abstract
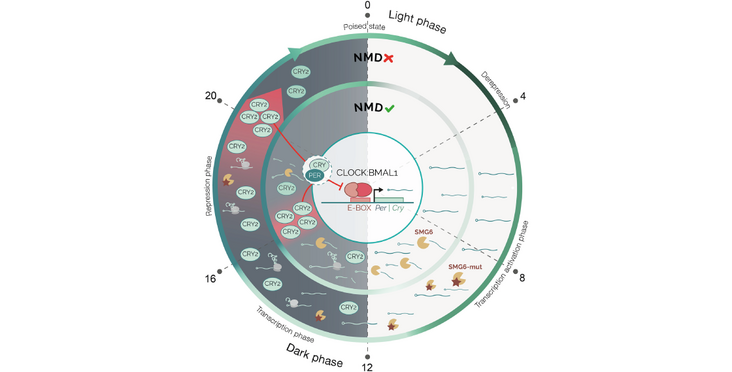
Nonsense-mediated messenger RNA (mRNA) decay (NMD) has been intensively studied as a surveillance pathway that degrades erroneous transcripts arising from mutations or RNA processing errors. While additional roles in physiological control of mRNA stability have emerged, possible functions in mammalian physiology in vivo remain unclear. Here, we created a conditional mouse allele that allows converting the NMD effector nuclease SMG6 from wild-type to nuclease domain-mutant protein. We find that NMD down-regulation affects the function of the circadian clock, a system known to require rapid mRNA turnover. Specifically, we uncover strong lengthening of free-running circadian periods for liver and fibroblast clocks and direct NMD regulation of Cry2 mRNA, encoding a key transcriptional repressor within the rhythm-generating feedback loop. Transcriptome-wide changes in daily mRNA accumulation patterns in the entrained liver, as well as an altered response to food entrainment, expand the known scope of NMD regulation in mammalian gene expression and physiology.
Read the Publication in Science Advances
Abstract, figure and title from Katsioudi et al (2023) Science Advances published under a CC BY-NC 4.0 license.