The Jinek and Kutay labs recently joined forces to gain functional, mechanistic and structural insights on the RNA helicase DHX37 and how it is activated by UTP14 during ribosome assembly. By determining the crystal structure of DHX37 bound to a single-stranded RNA substrate, the study provides critical insights into the RNA translocation mechanism of processive RNA helicases. Another key finding is that the ribosome biogenesis factor UTP14 interacts with DHX37 to enhance its RNA binding and stimulate its ATP-dependent RNA remodelling activity. This interaction serves as a key mechanism to activate DHX37 during a key step in the assembly of the small ribosomal subunit.
Abstract
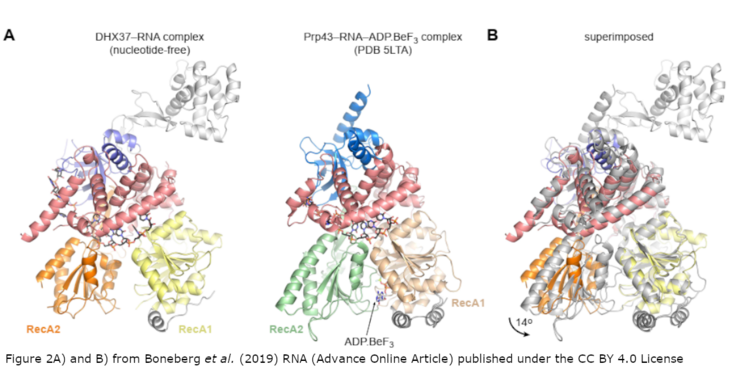
Eukaryotic ribosome biogenesis is a highly orchestrated process involving numerous assembly factors including ATP-dependent RNA helicases. The DEAH helicase DHX37 (Dhr1 in yeast) is activated by the ribosome biogenesis factor UTP14 to facilitate maturation of the small ribosomal subunit. We report the crystal structure of DHX37 in complex with single-stranded RNA, revealing a canonical DEAH ATPase/helicase architecture complemented by a structurally unique C-terminal domain. Structural comparisons of the nucleotide-free DHX37-RNA complex with DEAH helicases bound to RNA and ATP analogs reveal conformational changes resulting in a register shift in the bound RNA, suggesting a mechanism for ATP-dependent, 3'-5' RNA translocation. We further show that a conserved sequence motif in UTP14 interacts with and activates DHX37 by stimulating its ATPase activity and enhancing RNA binding. In turn, the C-terminal domain of DHX37 is required, but not sufficient, for interaction with UTP14 in vitro and is essential for ribosome biogenesis in vivo. Together, these results shed light on the mechanism of DHX37 and the function of UTP14 in controlling its recruitment and activity during ribosome biogenesis.
Read the Publication in RNA (Open Access)
Figure 2A) and B) and abstract from Boneberg et al. (2019) RNA published under the CC BY 4.0 licencse.