The Allain lab (IBC) and collaborators described in a "Science Advances" publication the unexpected mechanism of how an RNA-dependent negative feedback loop represses transcription by changing the chromatin state. Their findings shed light on yet another way of RNA-mediated gene regulation.
The transcriptional activator Mixed-lineage leukemia (MLL1) belongs to the HOX family and its fusions with other proteins constantly activate the transcription of target genes in certain leukemias. All fusion proteins lack three MLL1 PHD domains, including PHD3. Reinsertion of PHD3 into fusion proteins blocks their oncogenic function. The PHD3 domain binds methylated histone H3 lysin 4 (H3K4me3) and Cyclophilin 33 (Cyp33), the latter binding leading to transcriptional silencing by an unknown mechanism.
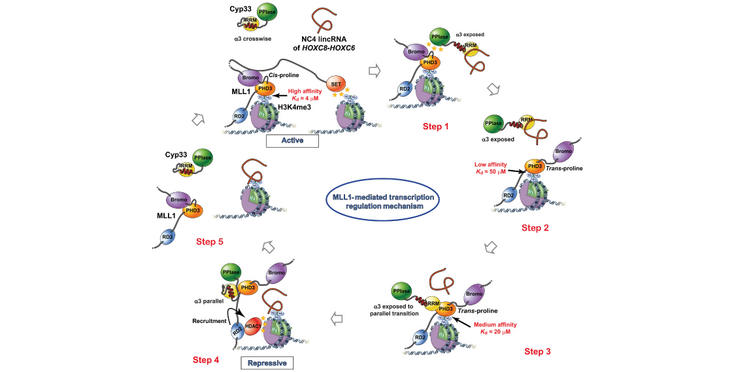
To shed light on the mechanism, the researchers solved five NMR structures complemented by biochemical and cellular assays and network modeling. They discovered a negative feedback loop mechanism to suppress MLL1-activated transcription based on RNA. In brief, MLL1 causes transcription of long intergenic long noncoding RNAs, which recruit Cyp33 via its RNA Recognition Motif (RRM) to the chromatin site where MLL1 is bound. The peptidylprolyl isomerase (PPI) domain of recruited Cyp33 then isomerizes a specific MLL1 proline, allowing the RNA-bound Cyp33 RRM to bind to the PHD3 domain releasing it from H3K4me3 with the help of the bound RNA. H3K4me3 can then be demethylated, leading to transcriptional repression of MLL1 target genes.
A key role in orchestrating this process resides in the Cyp33 conserved linker region between the RRM and the PPI domains, shown to be alpha-helical and adopting three different positions during the silencing process. The findings underscore Cyp33's fundamental role in repressing MLL1 transcription dysregulated in certain leukemias and revealed an unexpected novel mode of RNA-mediated gene expression regulation.
Read the Publication in Science Advances (Open Access)
Figure from Blatter et al. Sci Adv (2023) published under a CC BY 4.0 license.